Caloric restriction reverses aging: one cell at a time
Aging is known to cause functional decline throughout the whole body[1]. How can you study geroprotective interventions on the cellular and molecular levels across an entire organism? Single-cell transcriptomic atlasing has come to the rescue! In a tour-de-force, Ma et al.[2] dissected the effect of caloric restriction (CR) in response to aging by generating a transcriptomic atlas of over 210,000 individual cells and nuclei across nine rat tissues.
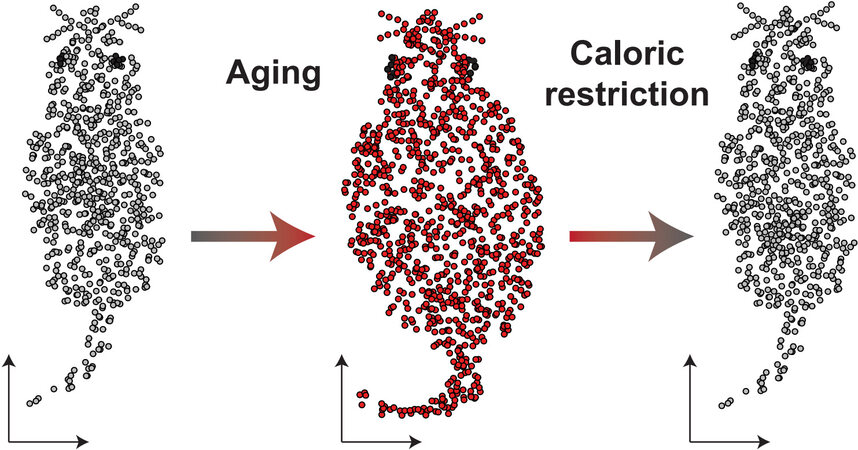
From an analytical perspective, the authors leveraged a powerful double contrast approach [Figure 1]. In the first contrast, young (Y-AL) and old (O-AL) ad libitum fed rats were compared to each other to characterize the impact of aging. In the second contrast, the O-AL rats were compared to old rats undergoing CR (O-CR) to characterize changes in response to CR.
Figure 1. Single-cell transcriptomics dissects impact of caloric restriction on aging at the molecular and cellular levels. Using a double contrasts analysis strategy the authors first defined alterations associated with normal aging and caloric restriction followed by comparison of both contrasts. This analysis facilitated the discovery of aging associated changes that were “rescued” upon caloric restriction.
This analytical strategy enabled the authors to disentangle alterations in response to aging from the effect of CR. Most critically, by comparison of these two contrasts, the authors were able to identify aging-associated alterations that were “rescued” by CR.
Analysis of cell type frequencies across both contrasts revealed an increase of immune cells in multiple tissues except bone marrow during aging. Interestingly, these increases were not observed when contrasting the O-AL with O-CR groups and thus were prevented by CR.
Next, the authors investigated alterations at the gene expression level within each cell type. Based on the two aforementioned contrasts, thousands of differentially expressed genes (DEGs) associated with aging and/or CR were identified. Integrative analysis of these two contrasts revealed many “rescue DEGs”, i.e., DEGs associated with aging that were partially rescued by CR. While the degree of rescue lacked quantification, the analysis revealed that a large fraction of transcriptional changes was rescued by CR. Only 1.8% of aging DEGs showed concordant changes in the CR contrast. For example, a gene increased with age and then increased even further upon CR. The small fraction of genes sharing this pattern may represent the noise in the system and, thus, suggest minimal side effects in rats.
Subsequently, the biological pathways underlying the gene expression changes were interpreted using Metascape[3]. The authors observed that upregulated rescue DEGs were involved in development, regeneration, the response to growth factors, extracellular matrix organization, and the response to corticosteroids. On the other hand, downregulated rescue DEGs were involved in inflammation, the innate immune response, the response to lipopolysaccharide, the response to interleukins, reactive oxygen species metabolic processes, and apoptotic signaling pathways.
When looking for genes that were altered constitutively across many cell types, the authors highlighted S100a9, S100a8, and Igkc, which were upregulated in more than 40 cell types during aging and downregulated by CR in more than 30 cell types. Moreover, Ybx1 was downregulated in more than 30 cell types during aging and upregulated by CR in more than 20 cell types. The authors selected Ybx1 to validate their findings by performing independent bulk RNA-seq experiments. Knockdown of Ybx1 in adipose-derived stem cells resulted in progressive loss of cell proliferative potential consistent with the exhaustion of this type of stem cells in vivo. These experiments validated the computationally derived findings and suggested that Ybx1 may represent a key molecular switch between the effects of aging and CR.
To elucidate the transcriptional regulators underlying the gene expression changes, an algorithm called SCENIC[4] was applied. SCENIC infers activity of transcriptional regulators at the cellular level and thus facilitates the study of cell type specific regulation. The analysis revealed rescued activity by CR for several transcription factors. The authors highlighted Cebpd and Cebpb because both transcription factors were commonly downregulated during aging and upregulated by CR in multiple tissues. Both of these transcription factors have previously been linked to inflammation and lipid metabolism and thus suggest interregulation by aging and CR.
The authors next used their single-cell RNA sequencing data to model cell-cell communication networks by mapping annotated receptor-ligand pairs across cell type marker genes using an algorithm called CellphoneDB[5]. Across several tissues, endothelial cells most prominently increased their interactions with other cell types during aging. However, these interactions were repressed upon CR, suggesting that endothelial cells may represent a target for aging intervention mediated by CR. However, a bias towards endothelial cells in the existing receptor-ligand annotations may give a technical reason as to why this cell type was so identified.
In conclusion, the authors generated a comprehensive whole-animal level single-cell rat atlas, which represents a valuable resource for the research community with interests in aging, caloric restriction, rat biology, and single-cell transcriptomics. Leveraging cutting-edge computational analyses, the effect of CR on aging was studied at the cell type frequency, gene expression, transcriptional regulation, and cell-cell communication levels. Taken together, the results demonstrate the rescue of several age-related phenotypic alterations in response to CR, providing a molecular basis for the geroprotective effects of CR.
DECLARATIONS
Authors’ contributionsThe author contributed solely to the article.
Availability of data and materialsNot applicable.
Financial support and sponsorshipNone.
Conflicts of interestThe author declared that there are no conflicts of interest.
Ethical approval and consent to participateNot applicable.
Consent for publicationNot applicable.
Copyright© The Author(s) 2021.
REFERENCES
1. Campisi J, Kapahi P, Lithgow GJ, Melov S, Newman JC, Verdin E. From discoveries in ageing research to therapeutics for healthy ageing. Nature 2019;571:183-92.
2. Ma S, Sun S, Geng L, et al. Caloric restriction reprograms the single-cell transcriptional landscape of Rattus norvegicus aging. Cell 2020;180:984-1001.e22.
3. Zhou Y, Zhou B, Pache L, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 2019;10:1523.
4. Aibar S, González-Blas CB, Moerman T, et al. SCENIC: single-cell regulatory network inference and clustering. Nat Methods 2017;14:1083-86.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Simon LM. Caloric restriction reverses aging: one cell at a time. J Cardiovasc Aging 2021;1:7. http://dx.doi.org/10.20517/jca.2021.12
AMA Style
Simon LM. Caloric restriction reverses aging: one cell at a time. The Journal of Cardiovascular Aging. 2021; 1(1): 7. http://dx.doi.org/10.20517/jca.2021.12
Chicago/Turabian Style
Simon, Lukas M.. 2021. "Caloric restriction reverses aging: one cell at a time" The Journal of Cardiovascular Aging. 1, no.1: 7. http://dx.doi.org/10.20517/jca.2021.12
ACS Style
Simon, LM. Caloric restriction reverses aging: one cell at a time. J. Cardiovasc. Aging. 2021, 1, 7. http://dx.doi.org/10.20517/jca.2021.12
About This Article
Copyright
Data & Comments
Data

 Cite This Article 9 clicks
Cite This Article 9 clicks











Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.